Hunter Dodson • April 18, 2024
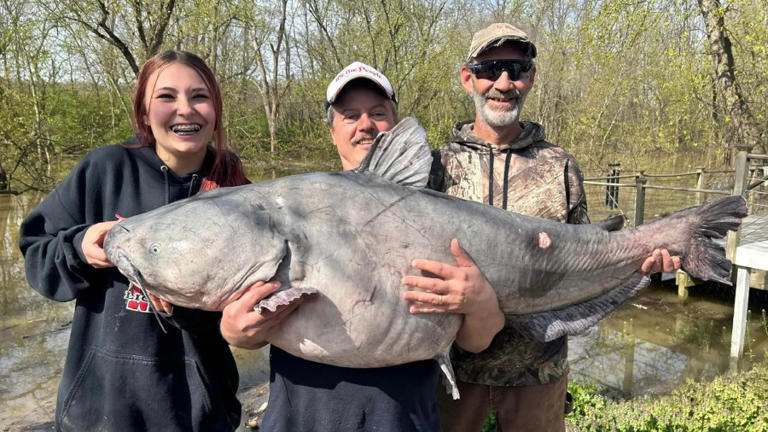
A teenage girl has recently broke the record for the largest fish caught in Ohio. On Sunday, April 7th, Jalynn Parker was doing some casual fishing...

April 16, 2024

April 7, 2024

April 2, 2024

Hunter Dodson • April 7, 2024
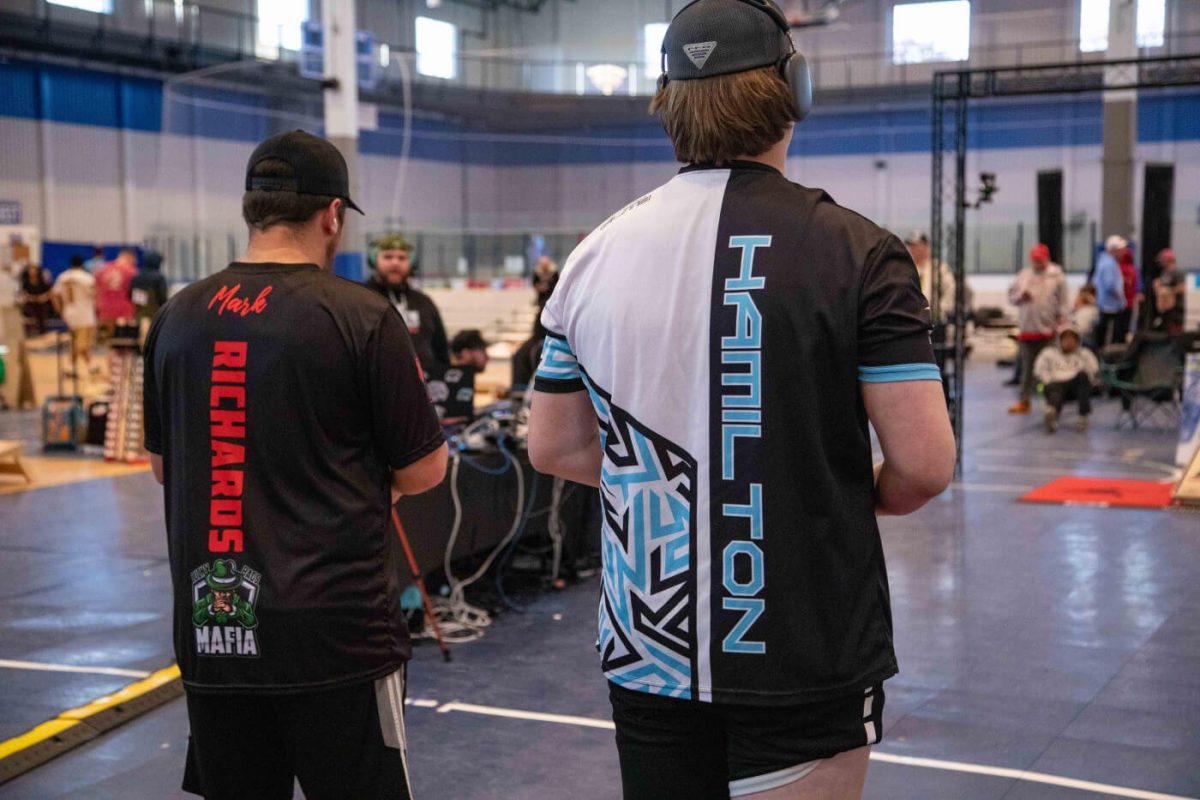
This past month, the American Cornhole League OPEN 11 took place at the Highlands Sports Complex in Triadelphia, West Virginia. The ACL OPEN...

December 7, 2023

December 7, 2023

November 4, 2023

October 12, 2023
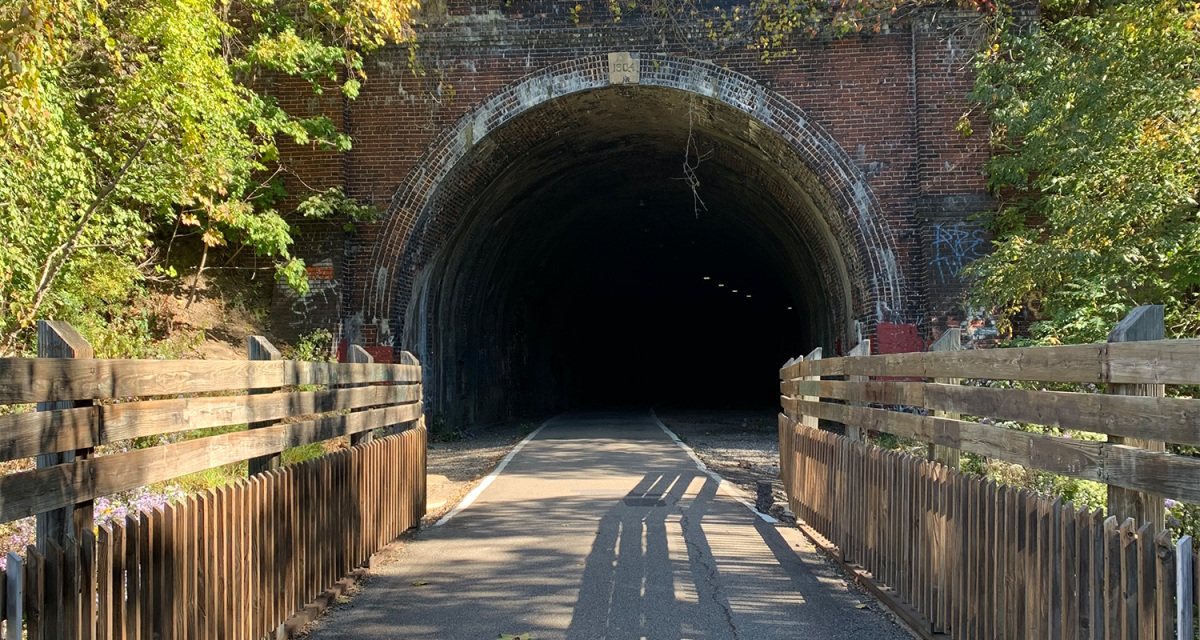

David Wiselka • April 8, 2024
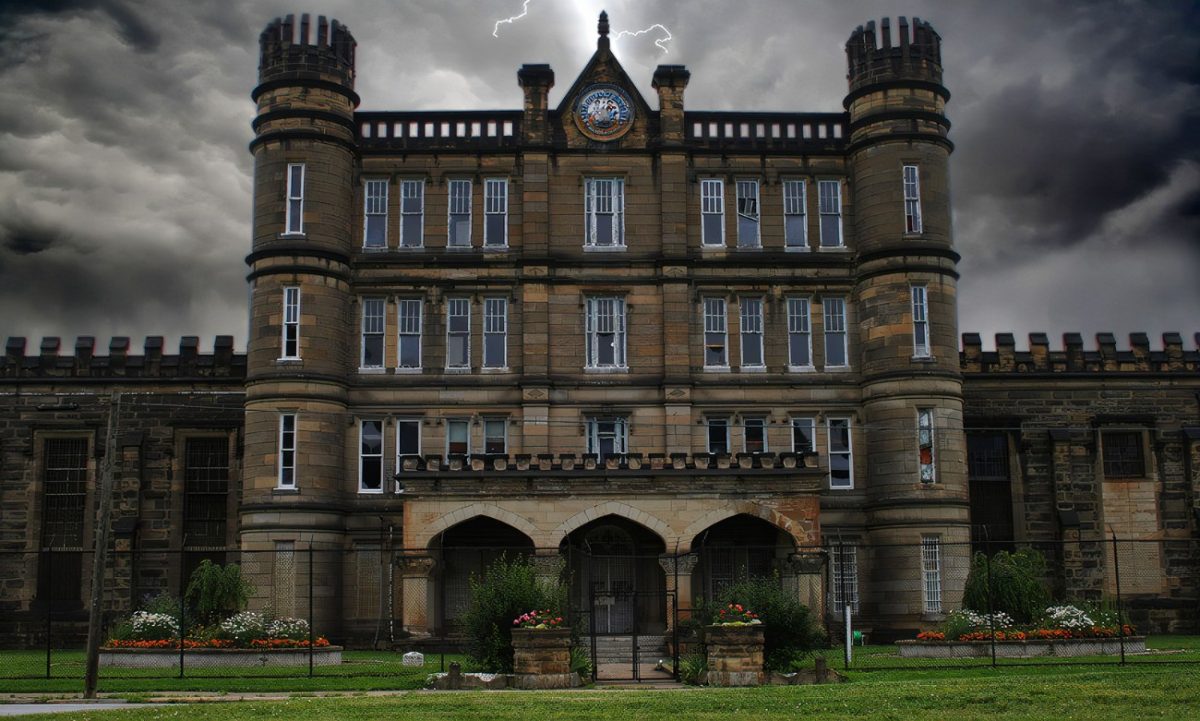
Placed in the heart of Wheeling, the Hempfield Tunnel, also known as "Tunnel Green," is a captivating and haunting structure that has stood the...

March 28, 2024
March 28, 2024

March 23, 2024

Finals week can be one of the most stressful weeks for college students. It is full of having to make sure all your assignments are completed...

October 2, 2023
October 21, 2022

October 3, 2022
Trending Stories







